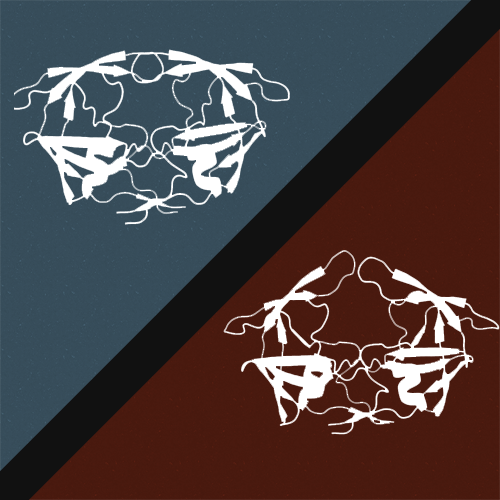
High-pressure NMR
High-pressure is a well-known perturbation method used to destabilize globular proteins and dissociate protein complexes or aggregates. In contrast to other perturbation methods such as heat or chemical denaturant, pressure affects locally the regions or domains of a protein containing internal cavities rather than destabilizing uniformly its structure.
In addition, pressure perturbation is generally reversible, which is essential for a proper thermodynamic characterization of a protein equilibrium. We use high-pressure perturbation combined with solution NMR spectroscopy to monitor at an atomic resolution the unfolding of monomeric proteins and the dissociation of protein complexes.
Luan M. Nguyen, Roche J. (2017) High-pressure NMR techniques for the study of protein dynamics, folding and aggregation. J. Magn. Reson. 277: 179-185
Roche J, Louis J.M, Bax A, Best R. (2015) Pressure-induced structural transition of mature HIV-1 Protease from a combined NMR/MD simulation approach. Proteins. 83: 2117-2123
Roche J, Dellarole M, Caro J.A, Norberto D.E, Garcia A.E, Garcia-Moreno B, Roumestand C, Royer C.A. (2013) "Effect of internal cavities on folding rates and routes revealed by real-time pressure-jump NMR spectroscopy". J. Am. Chem. Soc. 135: 14610-14618

Scaffold proteins and disordered peptides involved in neurological diseases
Intrinsically disordered proteins or peptides and scaffold proteins with long disordered regions are abundantly produced in eukaryotes and are known to be essential components of the cellular signaling machinery.
We are interested in the characterization of disordered peptides and scaffold proteins involved in neurodegenerative diseases (Alzheimer, Parkinson) and neurological disorders (schizophrenia, bipolar disorder). We use solution NMR, X-ray crystallography and fluorescence spectroscopy to characterize these proteins and understand their role and contribution to neurological dysfunctions.
Roche J, Shen Y, Jung Ho L, Jinfa Y, Bax A. (2016) Monomeric Aβ1-40 and Aβ1-42 peptides in solution adopt very similar ramachandran map distributions that closely resemble random coil. Biochemistry 55: 762-775
Roche J, Ying J, Bax A. (2015) “Accurate measurement of 3JHNHα couplings in small or disordered proteins from WATERGATE-optimized TROSY spectra”. J. Biomol. NMR 64: 1-7
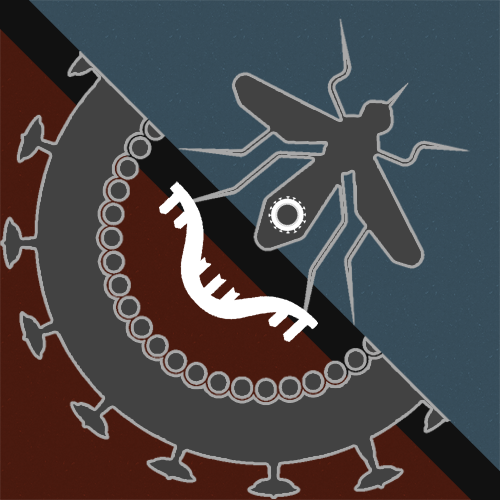
Evolution of viral proteins
The rapid evolution of viral genomes poses considerable challenges for the development of efficient anti-viral strategies. We seek to understand the mechanisms by which retroviruses (HIV) and flaviviruses (yellow fever, dengue and Zika virus) can adapt to new hosts, escape from the immune system and develop drug-resistance mutations, using a combination of bioinformatics analysis and biophysical methods (solution NMR and calorimetry).
Louis J.M, Roche J. (2016) Evolution under drug pressure remodels the folding free-energy landscape of mature HIV-1 protease. J. Mol. Biol. 428: 2780-2792

Protein folding
Most proteins fold into their lower energy conformation through a complex mechanism determined by the underlying free-energy landscape.
We seek to characterize the delicate equilibrium between the folded, unfolded and potential intermediate states of globular proteins, protein complexes and fibrils involved in human pathologies through a wide range of biophysical techniques including solution NMR, fluorescence spectroscopy and calorimetric methods.
We also use various computational techniques including enhanced molecular dynamics simulations and structure-based (“Go-model”) simulations.
Roche J, Dellarole M, Caro J.A, Guca E, Norberto D.E, Yang T, Garcia A.E, Roumestand C, Garcia-Moreno B, Royer C.A. (2012) "Remodeling of the folding free-energy landscape of staphylococcal nuclease by cavity-creating mutations". Biochemistry 51: 9535-9546
